A linear-encoding model explains the variability of the target morphology in regeneration
Paper Review
I’m always kicking myself for not writing more paper summaries/reviews/etc. They’re so helpful later on. So I’m gonna start! I’ll probably only write up stuff for ~10% of what I read, most of it won’t meet the threshold of interesting/important/cool. But when it does I’m gonna write up a post on it, which is mostly for myself, but hopefully also helpful for others!
Alright so this paper is titled: “A linear-encoding model explains the variability of the target morphology in regeneration” which is a bit of jargon for what seems like a really useful idea.
I think the basic idea is well said in an analogy they present in the paper. Ant colonies have all kinds of emergent complexity, if you zoom out to look at group behavior. But at an individual level, each ant is just following some simple, local rule like “follow the pheromone trail.” Most of biology is interested in understanding these simple, local rules which, when combined, results in all of the complexity of biological form and function. These rules are usually things like the interactions in gene regulatory networks (i.e., how do these genes interacting in this way give rise to phenotype X?) and this is the bulk of the focus in molecular and synthetic biology.
But, as the authors point out, if you want to manipulate colony behavior, it doesn’t seem obvious at all how one would change the individual rules to get some specific outcome. Say you wanted to build a chimney in the colony (which is apparently a thing for ants!) at coordinate X – it’s surely possible to change the individual, local rules to get this result, but it seems pretty difficult.
Similarly, with something like the Game of Life, one can easily see how a glider is formed from a feedforward application of the single cellular automata rule, but it’s much harder to reverse engineer this problem (i.e., going from glider to rule).
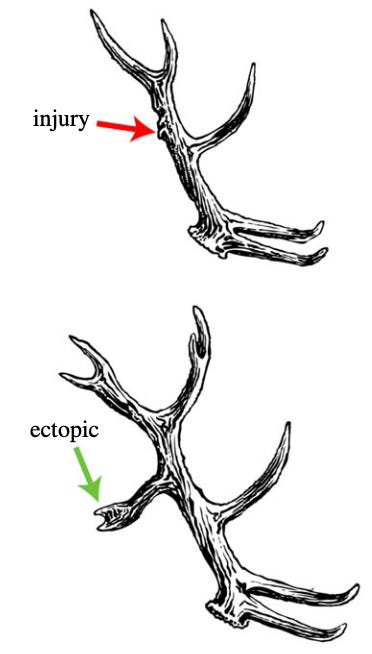
The authors claim that this reverse problem is what organisms are solving when they regenerate. Take deer, for instance. They have this amazing property that when their antlers are injured, they will grow a tine in the wounded area in the next season. (Schematic in the image below).
This seems an awful lot like building a chimney in the ant colony. There is some large-scale, anatomical change that the organism is trying to target.
And while it’s possible that this is engendered by simple, local rules – it’s a bit hard to imagine this is the most parsimonious way to do something like this. The antlers fall off, so you can’t just do some local processing at the wounded site, the wounded site isn’t there! The cells at the base of the antler need to “remember” for seasons, what the shape is that they are going to manifest. How this could be accomplished by something like the bottom-up rules in Game of Life seems incomprehensibly hard. Instead, the authors argue, there is a much more straightforward, top-down encoding of the “target morphology” somewhere in the organism. And it is this encoding which guides the cellular mechanics.
Before I get to what that encoding is, I want to list a few of the truly spectacular things that happen in biological regeneration. For one, if you completely scramble an embryo (like mix up the cells), it will rejigger itself into the correct form and create a perfectly normal adult animal. You can do a similar thing to frogs: mix up their face in development, and it will return to normal form (which requires completely bizarre movement – mouths and eyes moving around in ways that likely never happened in its evolutionary history). You can graft the tail of a salamander onto an amputated limb and the tail will slowly morph into an arm. Also, when you starve planaria (a small worm), they continuously adjust their organ sizes to be in the correct proportion as their cell numbers decrease.
It’s pretty incredible. The deer tines above are another example. One of my all time favorites, though, is that you can experimentally enlarge the cell sizes in newt kidney tubules, and as they get bigger fewer and fewer cells will get recruited to keep making the “right sized tubule.” If you make the cells huge, a single cell will wrap around to keep the size correct.
The authors go through three examples in detail: antler, planaria, and fiddler crab regeneration. We’ve already mostly been through the antler example, but one interesting thing to note is that when you anesthetize the animal, and injure the antler, the morphology doesn’t change in the next season, it just reproduces whatever antler structure it had the previous year. It seems likely that some electrical network (which anesthesia disrupts) is maintaining the target morphological encoding.
With planaria, you can amputate the tail and selectively change the cellular electrical communication and lo and behold a head will grow there instead, producing a two-headed worm. One thing to note here is that the pattern is long-term: when you cut the head off the two-headed worm in plain water, a head regrows. This is all without any direct modification to DNA. The authors point this out as a way in which biology’s typical mode of explanation fails: if a biologist were to find this worm in the wild (the two headed one), they would probably search for a speciation event. This would be misleading though, since the morphology was induced completely outside of any genetic modification.
Another thing to note is that you can trim planaria down quite a bit (the record is 279 cuts), and from all of these the same worm will regrow. This seems to imply that the target morphology is not stored in a one-to-one mapping (since the resulting worm is bigger), but in a compressed and distributed form throughout the worm.
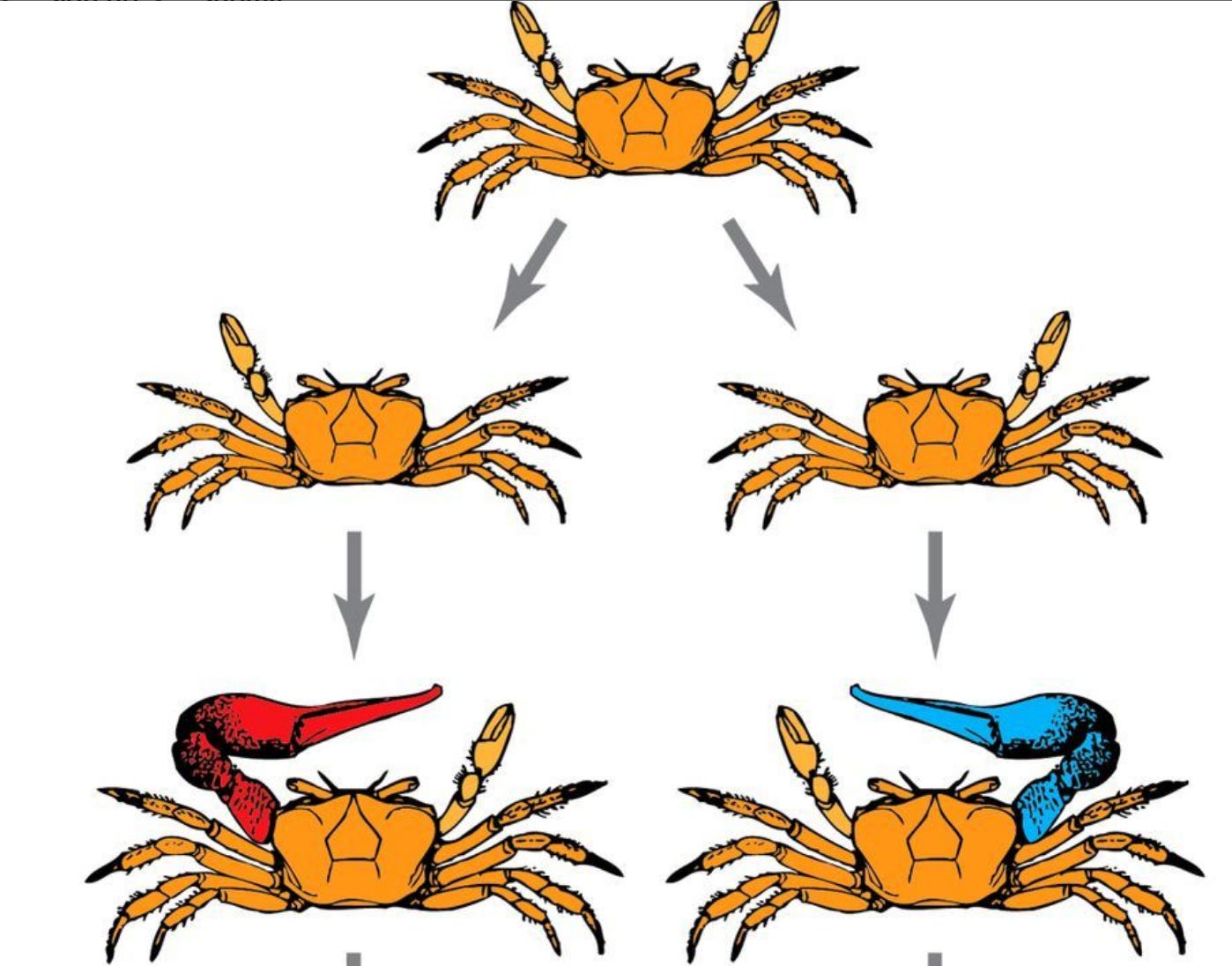
The fiddler crab also has variable morphology: sometime in its development, one of its claws will fall off (this always happens as a result of eating/fighting/etc). The side where it falls off first is the side which regrows a massive claw, and this asymmetry persists throughout its life (pic below). This is not genetically induced, instead it’s the result of random environmental effects (about half are right-handed, half are left, but you can’t read this out of the genome).
Okay so what the hell is this encoding? The authors claim that in order to do regeneration, organisms need to solve this inverse problem. They explain that you can think of typical biological development as being equivalent to something like G(m) = d, where G is an operator representing the physical events which reliably shape m, the parameters of the organism (genome), and d is the resulting morphology. Regeneration, then, is the opposite: going from d to m.
Going from d to m is much easier with linear encodings. A linear encoding is something which straightforwardly relates things in the input to things in the output. So, for instance, in Game of Life, a glider is not well related to anything in the rule that creates it, you can’t really read it out from the rule itself, you have to run it forward. But take something like ‘WWWWWBBBB’ – you could represent this as 5W4B (i.e., ‘five whites and four blacks’), a much shorter string, but which relates the input to the output in an exact and straightforward way. With linear encodings, the important feature is that working backwards is easy. It’s pretty hard to go from glider to rule, but extremely easy to go from 5W4B to WWWWWBBBB. Another type of linear encoding is just a literal map, where everything has a one to one correspondence.
Another important feature is that, with linear encodings, a small, local change to the input results in a small, local change to the output. This is not so with non-linear encodings, which are more chaotic in nature. If you’re familiar with cellular automata, you’ll know that each rule creates wildly different “worlds.” The same is true with most non-linear dynamics. They showcase this with a toy model of antler regeneration. The specifics don’t feel super important to me here (basically in the linear case they use turtle grammar, in the non-linear case they use an L-system which is recursive) – the result seems generally true and important.
As you can probably see, if you make a small change to the input at any of the branches in the linear encoding, a small, local, and predictable change happens in the ensuing antler structure. If you do a small change at any of the locations in the non-linear encoding you get these crazy, hard to predict antlers, none of which are biologically feasible.
So, the authors claim, organisms that regenerate (and probably most animals, really), utilize linear encodings to do so. These are going to be representations that are stored somewhere in the organism. They guess that they are typically more compressed than a map (i.e., a one-to-one thing), since planaria can regenerate from a tiny portion of their tissue. I suspect Michael Levin’s guess is that these maps are primarily bioelectrical, although the authors are agnostic on this point (maybe they’re bioelectric, maybe they’re genetic, either way they’re encoded linearly). Although there seems to be more evidence pointing towards the representations being electric in nature.
This is pretty different from the standard biological framework (I think – not a biologist), where everything is thought to be derived from simple rules acting on genes which engender large-scale form, sort of like how gliders pop out of the game of life.
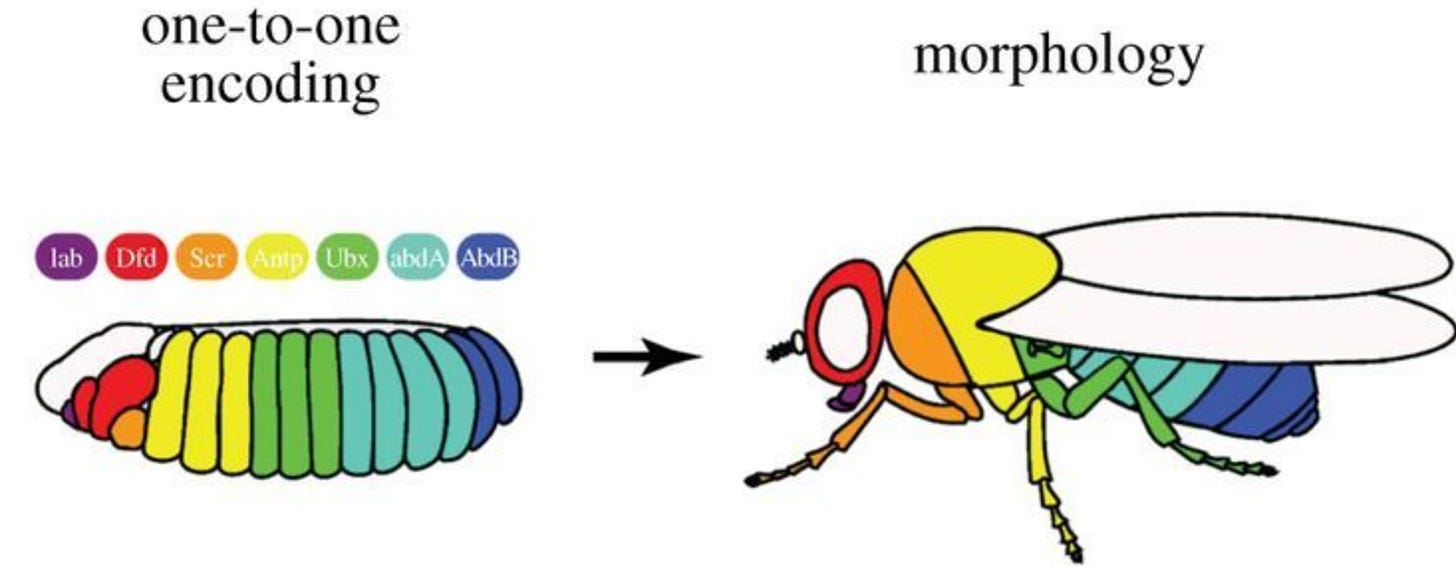
There’s evidence for these linear encodings, too. Apparently, there’s a two step process across many species development. First, you have some complex, emergent non-linear dynamic (like genetic networks). Then you get this linear encoding stage where everything nicely lines up with the final form the animal will take. You can see this in the schematic below for Drosophila (the second half of this process, where every stripe in the larvae one-to-one corresponds with some tissue in the adult fly).
If you change the genetic network responsible for the non-linear stage, really large, global changes will happen to the adult fly. If you alter the genes for going from the one-to-one encoding to the adult fly, small, local changes will result. This is also the case for tadpoles – if you change anything in the non-linear stage large-scale face deformities ensue. If you change the linear level (e.g., by altering the membrane potential of a small group of non-eye cells anywhere in the body), a whole ectopic eye will form (but just one eye, and locally). Both of these are in line with the toy antler model above – when you change the linear encoding, smaller, local change occurs.
And that’s basically it!
I think this paper is cool because it introduces an important concept that seems compelling to me: regenerative organisms (and probably most?) use something other than the feedforward, non-linear, DNA-as-blueprint model to guide their behavior. And whereas previously I thought that there had to be some abstract, global information guiding regenerative behavior, now I think the important thing is that it’s easy to move between desired state and current state (one-to-one mappings accomplish this which are not necessarily abstract or global). Whether or not that’s linear encodings per se doesn’t feel super relevant to me, but the general idea seems useful.
In some ways, this doesn’t seem radical at all: we clearly already assume something like this with brains. E.g., we think that the compressed representations instantiated in neural networks “causes” lower level cellular behavior like moving muscles or whatever.
But for some reason, biology usually shuns this mode of understanding. In this paper the authors mention that it’s because most biologists try to distance themselves from the theory of preformation (i.e., the idea that organisms develop from tiny little homunculi that resemble the adult form). I think biologists also shun anything that seems teleological because of the field's previous flirtation with vitalism. Now biologists want nothing to do with anything that seems top-down. The reigning paradigm is that organisms form from a DNA blueprint, more or less, like an instruction manual that interacts in complex ways with the environment, but a bottom-up process nonetheless, the culmination of simple, local interactions.
I think this view is importantly wrong (although of course also importantly right), in many of the ways the authors outline here. And I like this handle of linear vs non-linear. I especially liked their analogy of the ant colony, that seems like a really crisp and concise way of stating the problem.
But I have a few questions.
The first is that, when you amputate a planarian head and apply anesthesia, a new species head will regrow in proportion to where it is in the phylogenetic tree. Planaria have crazy DNA because of fission, so the species thing isn’t so weird here. The thing that is weird is that the head shape changes at all.
Levin’s explanation of this is that anesthesia disrupts the bioelectric network (which is where the morphological pattern is formed) and when it’s reactivated it tries to find its way back to normal, but there are different basins of attraction. So in proportion to how far away a head shape is in attraction-space from the current worm, you will probabilistically see it according to that distance.
Interestingly, and somewhat frighteningly, not all people “come back” from general anesthesia. There are reports of very long periods (on the order of years) of psychosis following anesthesia. Like their brains found a different basin of attraction in mental space. Yikes!
But when you anesthetized the deer, no morphological changes resulted in the antlers. My current best guess is that these encodings have something to do with bioelectric networks, but we don’t really know the connection yet, so maybe this just totally knocked it out, or maybe this basin is super reliable? But that with more precise editing we could get it to form weird new antler shapes with anesthesia? My second best guess of this is that they just didn’t do it enough times and that if you ran the experiment over and over you’d see changes.
My second question is whether or not all large, multicellular organisms have these linear encodings. The authors seem to imply that regeneration is a hard problem for non-linear encodings, but that straightforward development isn’t. It would be a pretty big bummer if only regenerative species had these linear encodings, because it would really limit our ability to extend therapeutic techniques based on this kind of work to humans. But I think, based on other things I’ve read, that Levin thinks this sort of thing would work in people to e.g., regenerate limbs, so – I’m confused! I suppose that in the fly example there is a one-to-one encoding although Drosophila isn’t regenerative. But then the question is, why? What’s the evolutionary pressure to have this if the organisms aren’t solving any hard problem with it?
Anyways, I think this line of work is really cool and seems incredibly worthwhile to investigate. If these linear encodings naturally occur in people, they seem like much more straightforward levers to press than our current approach of micromanaging genetic outcomes.