I started learning about biology a few months ago. Before knowing much about it I had confusions like: “what are transposons?? they can jump around the genome?? is this like a high tech epigenetic program?,”and “what does non-coding DNA do? why do complex animals have so much of it? does it set up online computers?”
And I wanted to know answers to questions like: “how do cells differentiate when they all have the same DNA?”, “how do cells overcome free energy gradients?”, “how is a human being made up out of a single cell,” “what’s the general framework biologists have and use to understand things?” and perhaps most importantly – “why do biologists dismiss Levin’s work?”
Michael Levin is a biologist I’m working with (well, trying to – I’m in the middle of writing grants to get a project going). He strikes me as one of the most important scientists currently doing work. He has a groundbreaking theory that could have huge implications for both medicine and for deconfusing concepts like “intelligence,” “agency,” and “goals.”
But for some reason barely anyone is talking about him. And I thought, well, I’m not a biologist, maybe there’s something really obvious that I’m missing which renders this pointless or dumb. I felt like this wasn’t going to be true, from what I did know, but I wanted to be more sure.
Now I feel much more confident that there is something real and interesting here. And I also learned a ton about biology and answered many of the above questions for myself. So here is all of that!
What I’ve Learned
I learned most of what I know from this fantastic book:
One of the most striking things about biology is that life is so fucking cool. You really gain an appreciation for just how complex and intricate every single part of an organism is after reading a cell biology textbook. It feels almost impossible – that something so complex could even happen, which is, of course, what billions of years of evolution looks like from the outside.
Cells are Molecular Machines (and Computers!)
Chemistry is foundational for life. In fact, many biologists simply call biology “applied chemistry.” Cells are like giant, churning, molecular machines. They are constantly breaking down and building up molecules – molecules to drive all kinds of processes within them.
Enzymes are proteins which specialize in this: they can increase the rate of chemical reactions by up to 10,000 fold. In effect, they lower the free energy gradient needed for a reaction (e.g., forming a bond) to occur. An example of how this actually works is with lysozyme (pictured below)– an antibiotic enzyme found in tears and saliva. Normally bacterial cell walls are not affected by water, and in the absence of an enzyme the random molecular collisions would almost never exceed the energetic threshold to cleave the wall. But when lysozyme latches onto the cell wall it reshapes it such that this reaction is much more likely to happen, thus cleaving it.
Another molecule that factors heavily into the creation and destruction of molecules is ATP. ATP is adenosine triphosphate. It’s very unstable (meaning that it “wants” to give up one of its phosphates – that's a negative free energy reaction). This can aid in creating molecules which would otherwise never be spontaneously made. For instance, glucose and fructose can join to make sucrose, but this would never randomly happen since it’s positive free energy. However, if you add ATP to the mix, you can get a coupled reaction where each step is negative free energy and which ultimately enables the creation of sucrose. This diagram was very helpful to me in understanding the concept:
The uncoupled reaction is nearly impossible, but once you add ATP, you can get this two step process (both steps are negative free energy overall), which ultimately results in the molecule you wanted.
This is only a two step process, but you can imagine chains that are arbitrarily long, and this is in fact what cells often do – they set up a series of enzymes to serially build up or break down particular molecules. These chains are what are generally referred to as “metabolism,” i.e., the cells way of taking energy from the environment and turning it into useful work.
And you might naturally ask, okay but how is ATP created? You can’t just create order out of nowhere, that goes against the second law of thermodynamics. Entropy must always increase. And indeed this is true. The high-level answer is that organisms consume order from their environment (e.g., through the ordered molecules in food), and break it down such that on net the entropy has increased, but it hasn’t increased in the ways that they “care about.”
One of the things that really struck me was how much of the molecular machinery was just really straightforwardly physical, as in, the specific shapes of molecules and proteins determine a lot of the activity in the cell. One example of a straightforwardly shape-based outcome is that saturated fats are made out of really long, straight strings of hydrocarbons whereas unsaturated fats have kinks:
You can imagine why saturated fats are solid at room temperature whereas unsaturated fats are liquid! The long chains stack much more easily than the kinky ones. Examples like this abound.
The properties of proteins (usually, the shape) are doing a lot of the work in cells. Proteins are just big, folded molecules (a sequence of amino acids). They’re the most abundant and complex molecules in the cell, and give rise to all the unique characteristics that vary from cell to cell. There are motor proteins, sensory proteins, signal proteins, storage proteins, there are proteins for everything!
Proteins also enable online computation in cells – many of them carry implicit if-then statements. For example, many enzymes have an inhibitory site – if a certain molecule attaches there, the entire shape of the enzyme transforms such that it stops working. In other words: “if inhibitory molecule → change state.” And just like you can get long strings of enzymes which catalyze a set of metabolic reactions – you can string together many proteins which, when activated, causes a domino effect of shape transformations to occur. These transformations drive things like cell movement, communication, and gene expression.
This is, for instance, how motor proteins “walk” along cytoskeletal filament. It’s a protein complex (also called a protein machine) which, through a series of shape transformations can cause, basically, walking behavior (pictured below). It’s unidirectional because it’s powered by ATP, and ATP only “wants” to go in one direction, i.e., to lose its phosphate. I found this very cool.
You can imagine how you can get all kinds of behavioral complexity from linking up different proteins in a series of reactions. There are things like cellular motors and pumps, too, in addition to all the metabolic reactions.
DNA and Epigenetics
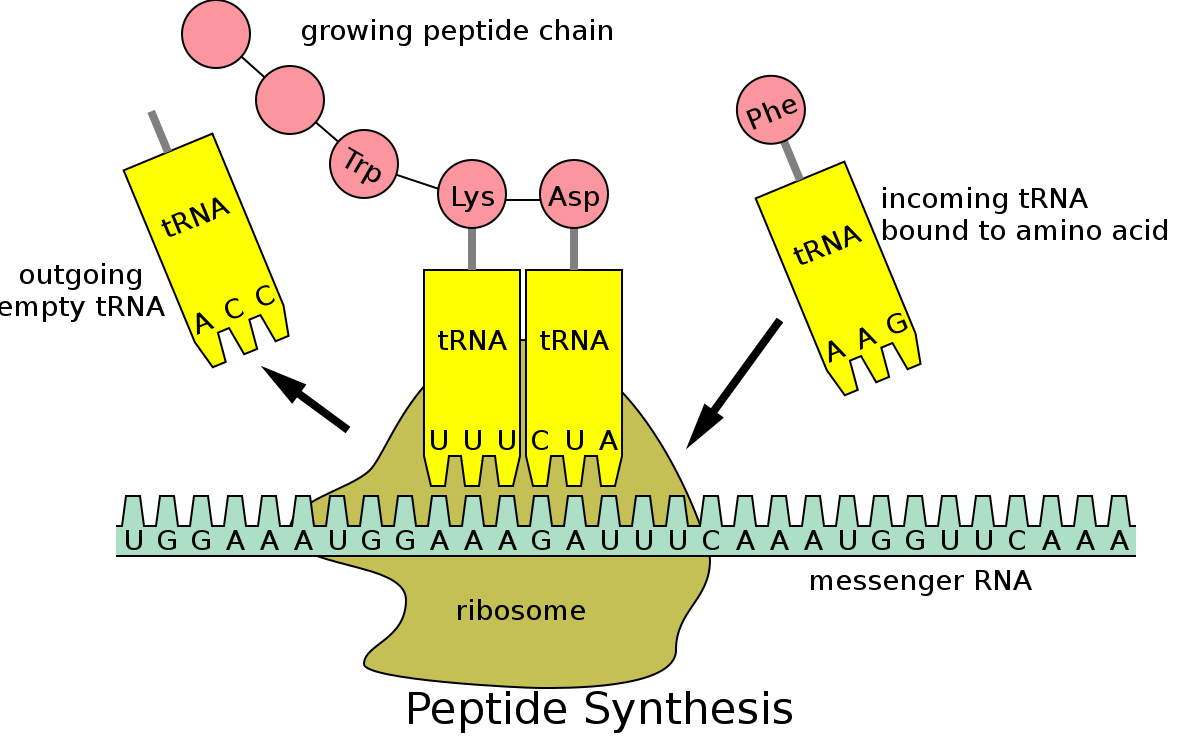
This leads us to DNA. DNA is the code which underlies protein formation in cells. Probably most people know this, but basically DNA is a long, double stranded helix which carries four nucleotides: A, T, G, and C. Triplets of these letters (e.g., AAG), correspond to specific amino acids (AAG, for instance, pairs to phenylalanine, or, Phe for short). They correspond in the sense that tRNA (t = transcription) molecules have the nucleotide sequence on one end and the amino acid attached to the other end (pictured below). A sequence of many of these triplets (e.g., a gene) corresponds to a specific amino acid sequence which assembles in the way shown below. This in turn corresponds to a protein.
So, DNA has genes, genes code for proteins, and proteins drive most of the cellular activity. So far so good. The only problem is, we know that not all genes are expressed (e.g., turned into proteins). In fact, in most mammalian cells, only half of the genes are ever expressed.
This has to be true because, e.g., every single cell has the same genome but their activity varies dramatically. A skin cell is doing a very different thing than a neuron. But what drives these changes? How does the cell “know” which genes to express?
The term for anything that affects gene expression is epigenetics (“epi” meaning “on” or “upon”). There are a lot of mechanisms to accomplish this (although we certainly don’t know all of them). I’ll highlight a few of them here, to give a feel for how it works.
Again, the physical structure of molecules is important here. DNA is extremely long, and hence has to be tightly coiled in order to fit into a cell. These coils, which are composed of DNA wound around proteins called histones, are what make up chromosomes. Their structure dictates what genes are more likely to be expressed: those wound up tightly can’t be accessed by RNA, those loosely coiled can be. One epigenetic role, then, is anything that acts on the histones, thereby changing the coiling, hence modulating genetic expression.
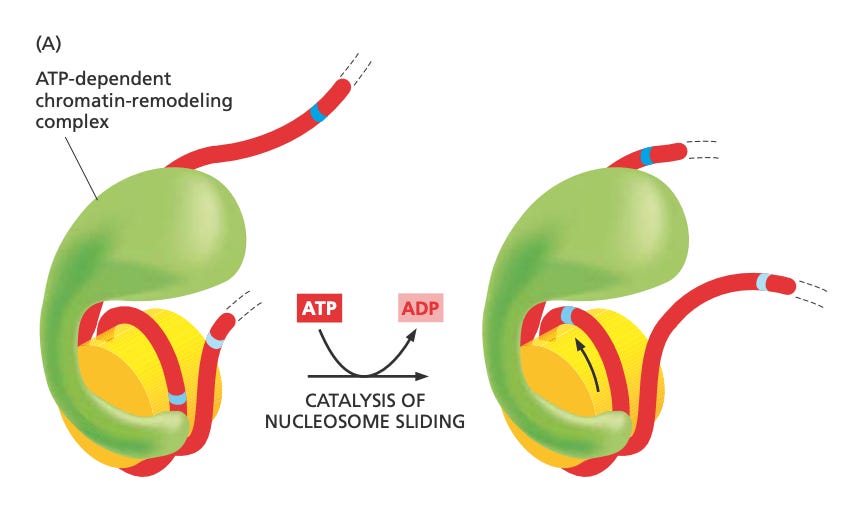
For instance, in the picture below, you have tightly bound chromatin (the yellow discs are histones and the red thread is DNA). Once it interacts with the green thing (an enzyme), ATP drives a unidirectional reaction of decondensing it. The spread out chromatin is easier to access, thereby enabling more genetic expression.
This is an up close picture of what the green protein complex is doing – pretty cool!
There are other ways too. Most (all?) genes have promoter sequences upstream of them – specific nucleotide sequences in the DNA with shape and electrical properties that make RNA polymerase bind to them especially strongly. This is usually what initiates gene expression. It can be turned off, though, basically by just clamping a protein onto the promoter site to inhibit RNA polymerase from binding there. For instance, a simple negative feedback loop might be: “if tryptophan concentration is too high, tryptophan will bind to a repressor protein, only in this condition will that repressor bind to the DNA promoter site, thus inhibiting the production of more tryptophan.”
That’s a simple circuit, but of course highly complex webs of if-thens exist which regulate genetic expression. They can be triggered or mediated by all kinds of things (chemical concentrations, extracellular or intracellular signals, protein transformations, etc.) If you’re familiar with the term “signaling cascades,” this is what it refers to. Long chains of command which ultimately results in genetic expression.
So, to answer myself from a few months ago: this is the solution to “how do cells differentiate if they all have the same DNA?” Essentially, a set of cues will trigger a little molecular computer to cause the “skin genes” to be expressed and the “neuron genes” to be repressed. One thing I’m still really curious about is how this cellular memory is passed on. When skin cells divide, they never spontaneously become neurons instead, which means that this epigenetic program (the one controlling the skin genes) is remembered. My book has so far not given me a much better understanding of this, but it seems incredibly interesting and I want to know more.
Non-coding DNA
Alright, so, DNA transcribes proteins. Surprisingly, though, only a very small percentage of it does this. The rest, some 98% of it in humans, was often called “junk DNA,” although today the popular term is “non-coding.” It’s just the part of the genome that we don’t think makes any proteins. Some parts of the non-coding DNA have been characterized as regulatory (i.e., epigenetic). Some parts are transposons. But most of it we just have no idea.
I’m currently unclear on what the regulatory mechanisms for non-coding DNA are, though. In the above examples, regulation was induced by “online proteins,” i.e., proteins which regulatory genes created such that when they, e.g., sense a chemical they down-regulate a certain other gene. But non-coding DNA doesn’t make any proteins, so what is it doing? One thing is promoter sequences, i.e., portions of DNA which aren’t transcribed but based on their properties can initiate genetic expression. These make up a very small percentage of non-coding DNA, though. But perhaps there are other things like this, too? Another thing might be that the stretches between coding DNA are actually useful because of the coiled structure, e.g., having a certain amount of space makes some genes more likely to be on the outside of the coil? This is just wild speculation, but I’m excited to get a fuller picture of this as I keep learning.
So at least part of the non-coding DNA is regulatory. We also know that a lot of it is composed of transposons. Transposons are “jumping genes.” They are the classic example of a selfish gene. They offer basically no advantage to the host, but they “try” to go on existing anyways, by copying and pasting themselves over and over into the genome (i.e., our genomes literally get longer as we age). This is obviously bad for the organism, since DNA damage is not great and you can disrupt a gene if you copy and paste into the middle of one. So, we have regulatory mechanisms to repress this kind of activity, but it sometimes happens anyways.
That’s mostly where I’m at now. I feel like I understand biology way better than I did a few months ago. My high-level takeaway for how biologists understand life is like: “There’s a series of chemical reactions mediated by proteins (made by DNA) that drive almost all cell functions. They can add up to do really complicated things like walking or forming really long, complex molecules. Proteins also function sort of like a computer in that they set up a ton of if-then statements that lock into each other to, e.g., use cues from the environment to trigger certain events (gene expression, expelling proteins outside of the cell, etc.). These can combine in really intricate, complex ways to form, e.g., the entire metabolic system.”
Miscellany
There were also a few interesting things that didn’t quite fit into the above narrative so I’ll add them as miscellany here:
Miscellany 1: Aging
Transposons are a (good) guess about what might mediate aging in humans. Typically, when there is damage to DNA, mitochondria go to a lower energy state (as more resources are focused on repairing damage), but they also release free radicals (why?? I don’t know, seems pretty bad, probably an unavoidable side effect?). Those free radicals then go on to cause more DNA damage. When cells are healthy, this doesn’t reach a positive feedback loop and the cell quickly returns to normal. But when damage has accumulated, a certain threshold is reached where this does positive feedback loop and results in senescence.
Transposons are one form of DNA damage. And as they grow in number, the likelihood of them replicating gets larger, too. So over time your DNA damage risk accelerates. When DNA is damaged, the cell redirects energy away from repressing transposons to fixing the damaged DNA. So, occasionally when you get a sunburn or whatever, transposons will have a chance to replicate. Usually this is fine, but, as mentioned above, after a certain threshold this will positive feedback loop (with transposons causing damage, leading to mitochondria sending out more free radicals thus causing more damage, and so on) until senescence. And it gets worse when your stem cells have accumulated transposons – this means that even your “new” cells come carrying a lot of potential for damage, and hence more propensity to senesce.
Miscellany 2: Development
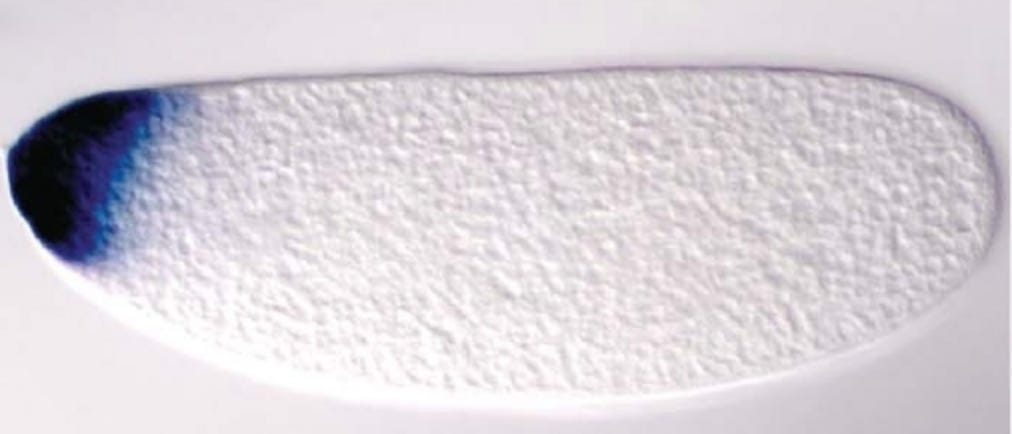
There were all these questions I had about how development works. Like, in order for a heart to develop on one side of an organism the cells in that area need to “know” that they’re on the left side – how do they know that? Well, I’m not sure about humans, but at least in flies, spatial axes are introduced via morphogen gradients. In other words, at a very early stage in embryo the top of the fly has more of a certain protein than the bottom (pictured below).
But then the obvious next question is: why do those cells have more of that morphogen?? It seems like you can just keep passing the buck. Then, I learned something that I thought was pretty interesting. There is what’s called a maternal to zygotic transition, which means that initially, the zygote (single-celled, fertilized egg), gets cues from the mother environment. Basically, the very early embryo does not do any of its own protein transcription, i.e., it doesn’t use its own DNA – the mother sends her RNA into the cell to kickstart the process. And there’s a transition in all animals wherein the embryo goes from relying entirely on the mothers RNA to its own DNA.
So, this is an answer to how location cues might be introduced: the mother sends in RNA to the top two cells in a four celled embryo saying “express these morphogens because you’re the top.” Of course, you could pass the buck one more time – how does she know? And I think at that point you’re just asking how cells initially discovered and propagated spatial cues to begin with, which can probably be explained by, e.g., whatever resulted in E. coli recognizing and following chemical gradients.
Michael Levin
But okay, to come full circle – how does any of this relate to Michael Levin? Well, Levin claims that organisms use software to direct their anatomy. In other words, in a similar way to how brains use electricity to navigate 3D space, Levin thinks that tissues use bioelectricity to navigate morphological space. Most of the above picture of biology is at the assembly code level: it’s working with the lowest-level parts (e.g., proteins, genes) and trying to figure out how they build up into these complex systems. It’s sort of like trying to understand a neural network from looking at the 1’s and 0’s and the AND/OR gates. Possible, but exceedingly difficult and inefficient.
Levin thinks that organisms have discovered ways to represent their anatomy at a higher level of abstraction in order to mediate and maintain macroscopic order. In particular, he thinks bioelectric maps tell cells when to e.g., stop growing, build new antlers, grow new limbs, what sorts of heads to build, etc.
So why do biologists disagree with Levin? I’ve talked with a few biologists and I’ve read some of their arguments. I think by and large their response rhymes a lot with the initial reactions to Darwin’s work on natural selection: “what’s true in them is old and what’s new in them is false.” Basically, biologists are like, “well, yeah, I mean we’ve known for a long time that electrical signals are cues for cells to do cell stuff, but they’re not special or anything. They’re just one of hundreds of other cues (like hormones, toxins, morphogens, etc) that can affect the cell mechanics.” Essentially, what’s true is old – that bioelectricity is important for cellular coordination, and what’s new is false – that it’s anything more special than that.
I expect that this is wrong. I could go into all my arguments here but most of this is coming up in the massive manifesto that I’ve been writing for the last eighty years and will totally, actually, for real publish soon. But basically, I feel pretty confident now that the field’s dismissiveness of Levin’s work has more to do with paradigmatic overshadowing than it has to do with the usefulness and truth of his ideas. And to be clear, no one is saying that his empirical work is wrong. It’s not like he’s a total outcast with no funding. It’s just that, for the sort of truly groundbreaking idea that this strikes me as, he’s getting relatively little attention.
Well I suppose I can’t resist giving a taste for why I suspect this is wrong (more details forthcoming). For one, regenerative species are capable of altering their morphology in real-time (e.g., deer can grow new antler structures over the course of their lifetime if they’re injured, pictured below).
In other words, they figured out how to grow a new, specific anatomy. So clearly the anatomical structure is not fully contained within the DNA. Where is it, then? The usual answer is epigenetics: somehow some long chain of if-thens results in the novel antler. But, remember, that antlers fall off. So, we are claiming that some non-local deer cells (at the base of the antler or something) remember more-or-less exactly what shape to manifest many months down the line and many miles away (from the cells point of view), relying only on these extremely low-level events. Maybe. But I’m not convinced.
Again, explaining this with the current biological picture is like trying to say how AlphaZero chose a new move in Go by consulting the CPU. Possible, and of course, ultimately this is what’s happening, but incredibly inefficient. And perhaps it is massively inefficient not just for us, but for the animal too– perhaps they have figured out more parsimonious, more abstract ways to represent the new antler than through the combinatorial nightmare of protein chains.
Secondly, we know that one biological structure has these properties: namely, brains. When we think of brains we generally consider their information content (or at least, their relevant content) to be at the level of neural firing patterns rather than, e.g., proteins making sure the neural cytoskeletons remain intact. The proteins are necessary, but they aren’t really what we care about. If we wanted to make big changes to someone’s behavior, we’re not going to poke a cytoskeletal protein, we’re going to try to change variables that are upstream, causal, and simple – things that affect the firing patterns, things like serotonin, or TMS, or even just plain therapy. The level of abstraction matters a lot when dealing with systems that span many orders of magnitude, and the field of biology is only carving out one or two of these levels, when life itself has figured out how to operate on all of them.
So, we already attribute these properties to brains which are, may I remind you, composed of cells just like any other section of your body. Brains also came onto the scene much later than other organs and systems – it seems entirely possible, suggestive even, that other electrical mechanisms for coordinating high-level, macroscopic outcomes were developed before organisms developed neurons to navigate 3D behavior.
Biology is very used to, and comfortable with, understanding its subject at one level of abstraction and it treats any other level skeptically. Levin’s work borders too close to preformation, is a breath away from vitalism, at least in the minds of biologists. Biologists deal with chemicals! Physics! From the outside it seems like Levin is proposing that teleology is in fact real – that organisms do have goals, real, high-level, abstract goals, that they optimize for. And that’s just, well, while not actually inconsistent with the current framework, it really smells of heresy.
I think these are very reasonable heuristics. I also think they’re wrong, here. Biology has gotten a lot of stuff right, and I’m not even claiming they’ve gotten anything “wrong,” per se, just that they’re using the wrong lens to look at macroscopic biological structure. And I think Levin is using the right one. He’s onto something incredibly interesting and groundbreaking, here, and I think the rest of the world should pay attention to it.